Optimización Bayesiana de hiperparámetros#
!pip install -q deepxde
[notice] A new release of pip is available: 26.0.1 -> 26.1
[notice] To update, run: pip install --upgrade pip
import os
# necesario para obligar a deepxde a usar el framework de pytorch.
os.environ['DDE_BACKEND'] = "pytorch"
import deepxde as dde
from matplotlib import pyplot as plt
import numpy as np
import skopt
from skopt import gp_minimize
from skopt.plots import plot_convergence, plot_objective
from skopt.space import Real, Categorical, Integer
from skopt.utils import use_named_args
import torch
torch.set_default_device("cpu")
if dde.backend.backend_name == "pytorch":
sin = dde.backend.pytorch.sin
pi = torch.pi
elif dde.backend.backend_name == "paddle":
sin = dde.backend.paddle.sin
else:
from deepxde.backend import tf
sin = tf.sin
pi = tf.constant(np.pi, dtype=tf.float32)
Using backend: pytorch
Other supported backends: tensorflow.compat.v1, tensorflow, jax, paddle.
paddle supports more examples now and is recommended.
Introducción#
En esta notebook vamos a resolver el problema de inversión para la ecuación de Poisson 1D en donde no conocemos el campo de fuerza, i.e.
La función pde(x, y) define el residuo físico que la red debe minimizar. Para eso calcula la segunda derivada de la salida respecto de la entrada, \(u''(x)\), y arma el residuo de la ecuación.
La función sol_exacta(x) no se usa para entrenar directamente toda la solución, sino como referencia para:
generar algunos datos observados,
evaluar el error de prueba,
comparar la solución aproximada con la exacta al final.
def pde(x, y):
u, q = y[:, 0:1], y[:,1:2] #ambas u y q son desconocidas
dy_xx = dde.grad.hessian(y, x, component=0, i=0, j=0)
return dy_xx - q
def sol_exacta(x):
return np.sin(np.pi * x)
Condiciones de contorno y datos observados#
Acá se construyen las restricciones que acompañan a la ecuación, exactamente como hemos hecho antes:
contornoidentifica si un punto pertenece al borde del dominio.func_contornofija el valor de la solución en el contorno, en este caso \(u=0\).x_trainyy_trainagregan puntos observados dentro del dominio, lo que ayuda a guiar el entrenamiento de la PINN.
# auxiliar para los puntos en el contorno
def contorno(x, en_contorno):
return en_contorno
# función auxiliar para los valores en el contorno
def func_contorno(x):
return 0
Supongamos que tenemos solo los datos de entrenamiento en puntos equiespaciados entre -1 y 0.
x_train = np.linspace(-1, 0, 10).reshape(-1, 1)
y_train = sol_exacta(x_train)
geom = dde.geometry.Interval(-1, 1)
observe_u = dde.icbc.PointSetBC(x_train, y_train, component=0)
bc = dde.DirichletBC(geom, func_contorno, contorno)
Construcción del modelo PINN#
La función create_model(config) recibe una configuración de hiperparámetros y arma una PINN completa. En esta etapa se definen:
el objeto
data, que contiene la ecuación diferencial, las condiciones y la discretización del dominio;la arquitectura de la red neuronal totalmente conectada;
el optimizador Adam y la métrica de error relativo \(L^2\).
Esto es importante porque la optimización bayesiana va a probar muchas configuraciones distintas, y en cada intento necesita reconstruir el modelo con otros hiperparámetros.
def create_model(config):
learning_rate, num_dense_layers, num_dense_nodes, activation = config
data = dde.data.PDE(
geom,
pde,
[observe_u, bc],
num_domain=16,
num_boundary=2,
solution=sol_exacta,
# anchors=x_train,
num_test=100,
)
net = dde.maps.FNN(
[1] + [num_dense_nodes] * num_dense_layers + [2],
activation,
"Glorot uniform",
)
model = dde.Model(data, net)
model.compile("adam", lr=learning_rate, metrics=["l2 relative error"])
return model, data
Entrenamiento y función de costo#
train_model ejecuta el entrenamiento durante un número fijo de iteraciones y recupera el historial de pérdidas. El valor que finalmente se devuelve como error es el mínimo de la pérdida de prueba.
Ese escalar resume qué tan bien funcionó una configuración dada y será la cantidad que la optimización bayesiana intentará minimizar.
def train_model(model, config, iterations=4000):
losshistory, train_state = model.train(iterations=iterations)
train = np.array(losshistory.loss_train).sum(axis=1).ravel()
test = np.array(losshistory.loss_test).sum(axis=1).ravel()
metric = np.array(losshistory.metrics_test).sum(axis=1).ravel()
error = test.min()
return error, losshistory, train_state
Espacio de búsqueda de hiperparámetros#
En esta sección se define qué aspectos del modelo se van a optimizar:
learning_rate: tasa de aprendizaje del optimizador.num_dense_layers: cantidad de capas ocultas.num_dense_nodes: cantidad de neuronas por capa.activation: función de activación.
También se fija default_parameters, que sirve como punto inicial conocido para arrancar la búsqueda. Este espacio mezcla variables continuas, enteras y categóricas, algo que skopt maneja de forma directa.
n_calls = 15
dim_learning_rate = Real(low=1e-4, high=5e-2, name="learning_rate", prior="log-uniform")
dim_num_dense_layers = Integer(low=1, high=6, name="num_dense_layers")
dim_num_dense_nodes = Integer(low=5, high=50, name="num_dense_nodes")
dim_activation = Categorical(categories=["sin", "sigmoid", "tanh"], name="activation")
dimensions = [
dim_learning_rate,
dim_num_dense_layers,
dim_num_dense_nodes,
dim_activation,
]
default_parameters = [1e-3, 4, 50, "sin"]
Función objetivo para la optimización bayesiana#
La función fitness(...) es la interfaz entre la PINN y el optimizador bayesiano. Para cada conjunto de hiperparámetros:
arma una configuración,
crea el modelo correspondiente,
lo entrena,
devuelve el error obtenido.
@use_named_args permite que skopt pase los parámetros en forma clara según los nombres definidos en dimensions. Además, si aparece un NaN, se reemplaza por una penalización grande para que esa configuración quede descartada por el algoritmo.
@use_named_args(dimensions=dimensions)
def fitness(learning_rate, num_dense_layers, num_dense_nodes, activation):
config = [learning_rate, num_dense_layers, num_dense_nodes, activation]
global ITERATION
print(ITERATION, "it number")
# Print the hyper-parameters.
print("learning rate: {0:.1e}".format(learning_rate))
print("num_dense_layers:", num_dense_layers)
print("num_dense_nodes:", num_dense_nodes)
print("activation:", activation)
print()
# Create the neural network with these hyper-parameters.
model, _ = create_model(config)
# possibility to change where we save
error, _, _ = train_model(model, config)
# print(accuracy, 'accuracy is')
if np.isnan(error):
error = 10**5
ITERATION += 1
return error
Búsqueda bayesiana de la mejor configuración#
gp_minimize implementa la optimización bayesiana modelando la función objetivo con un proceso gaussiano. En cada iteración usa la información acumulada de evaluaciones anteriores para decidir qué combinación probar después.
La función de adquisición EI (Expected Improvement) favorece puntos que prometen mejorar el mejor resultado actual, equilibrando exploración y explotación. Al terminar, search_result.x contiene los mejores hiperparámetros encontrados y search_result.fun el mejor error asociado.
ITERATION = 0
search_result = gp_minimize(
func=fitness,
dimensions=dimensions,
acq_func="EI", # Expected Improvement.
n_calls=n_calls,
x0=default_parameters,
random_state=1234,
)
best_params = search_result.x
best_score = search_result.fun
0 it number
learning rate: 1.0e-03
num_dense_layers: 4
num_dense_nodes: 50
activation: sin
Compiling model...
'compile' took 0.908025 s
Training model...
Step Train loss Test loss Test metric
0 [2.37e-02, 4.32e-01, 9.18e-04] [1.93e-02, 4.32e-01, 9.18e-04] [1.53e+00]
1000 [4.88e-04, 9.71e-04, 2.84e-06] [4.01e-04, 9.71e-04, 2.84e-06] [1.07e+01]
2000 [8.96e-05, 4.86e-04, 8.29e-06] [7.25e-05, 4.86e-04, 8.29e-06] [1.08e+01]
3000 [4.57e-05, 3.36e-04, 1.80e-05] [4.96e-05, 3.36e-04, 1.80e-05] [1.09e+01]
4000 [3.00e-05, 2.19e-04, 1.16e-05] [3.56e-05, 2.19e-04, 1.16e-05] [1.10e+01]
Best model at step 4000:
train loss: 2.60e-04
test loss: 2.66e-04
test metric: [1.10e+01]
'train' took 11.701705 s
1 it number
learning rate: 2.2e-03
num_dense_layers: 5
num_dense_nodes: 33
activation: tanh
Compiling model...
'compile' took 0.000136 s
Training model...
Step Train loss Test loss Test metric
0 [2.65e-01, 2.04e-01, 3.85e-01] [2.59e-01, 2.04e-01, 3.85e-01] [1.11e+00]
1000 [2.77e-04, 3.82e-03, 1.56e-05] [4.36e-04, 3.82e-03, 1.56e-05] [9.25e+00]
2000 [6.28e-05, 1.79e-03, 4.10e-05] [6.61e-04, 1.79e-03, 4.10e-05] [1.01e+01]
3000 [3.48e-05, 1.55e-03, 2.53e-05] [1.97e-04, 1.55e-03, 2.53e-05] [1.02e+01]
4000 [1.47e-04, 1.37e-03, 2.74e-05] [2.48e-04, 1.37e-03, 2.74e-05] [1.03e+01]
Best model at step 4000:
train loss: 1.54e-03
test loss: 1.64e-03
test metric: [1.03e+01]
'train' took 12.550742 s
2 it number
learning rate: 2.1e-02
num_dense_layers: 2
num_dense_nodes: 14
activation: tanh
Compiling model...
'compile' took 0.000301 s
Training model...
Step Train loss Test loss Test metric
0 [6.99e-02, 7.25e-01, 1.28e-01] [6.30e-02, 7.25e-01, 1.28e-01] [1.66e+00]
1000 [1.76e-04, 1.84e-03, 7.42e-05] [2.14e-04, 1.84e-03, 7.42e-05] [1.04e+01]
2000 [1.87e-02, 3.01e-03, 3.88e-03] [1.82e-02, 3.01e-03, 3.88e-03] [1.06e+01]
3000 [6.64e-03, 1.77e-03, 2.98e-03] [5.75e-03, 1.77e-03, 2.98e-03] [1.06e+01]
4000 [6.32e-05, 8.01e-04, 3.27e-05] [1.20e-04, 8.01e-04, 3.27e-05] [1.07e+01]
Best model at step 4000:
train loss: 8.97e-04
test loss: 9.53e-04
test metric: [1.07e+01]
'train' took 7.750267 s
3 it number
learning rate: 2.7e-04
num_dense_layers: 2
num_dense_nodes: 6
activation: sigmoid
Compiling model...
'compile' took 0.000115 s
Training model...
Step Train loss Test loss Test metric
0 [4.69e+00, 1.12e+00, 1.90e-01] [4.70e+00, 1.12e+00, 1.90e-01] [3.53e+00]
1000 [6.14e-01, 2.60e-01, 3.56e-02] [6.12e-01, 2.60e-01, 3.56e-02] [1.86e+00]
2000 [7.06e-02, 2.09e-01, 6.88e-02] [7.04e-02, 2.09e-01, 6.88e-02] [1.50e+00]
3000 [4.73e-03, 2.00e-01, 7.48e-02] [4.77e-03, 2.00e-01, 7.48e-02] [1.44e+00]
4000 [5.98e-04, 1.96e-01, 7.55e-02] [5.79e-04, 1.96e-01, 7.55e-02] [1.42e+00]
Best model at step 4000:
train loss: 2.72e-01
test loss: 2.72e-01
test metric: [1.42e+00]
'train' took 7.692979 s
4 it number
learning rate: 7.8e-04
num_dense_layers: 5
num_dense_nodes: 9
activation: sin
Compiling model...
'compile' took 0.000137 s
Training model...
Step Train loss Test loss Test metric
0 [1.56e-01, 5.04e-01, 7.02e-03] [1.34e-01, 5.04e-01, 7.02e-03] [1.77e+00]
1000 [2.96e-03, 2.68e-02, 1.82e-03] [2.57e-03, 2.68e-02, 1.82e-03] [6.73e+00]
2000 [4.18e-04, 1.52e-03, 5.18e-06] [4.72e-04, 1.52e-03, 5.18e-06] [1.01e+01]
3000 [7.49e-04, 8.52e-04, 5.37e-05] [7.19e-04, 8.52e-04, 5.37e-05] [1.06e+01]
4000 [1.45e-04, 7.38e-04, 4.63e-05] [1.87e-04, 7.38e-04, 4.63e-05] [1.07e+01]
Best model at step 4000:
train loss: 9.29e-04
test loss: 9.71e-04
test metric: [1.07e+01]
'train' took 12.374217 s
5 it number
learning rate: 1.6e-03
num_dense_layers: 1
num_dense_nodes: 18
activation: sin
Compiling model...
'compile' took 0.000101 s
Training model...
Step Train loss Test loss Test metric
0 [1.86e-01, 7.62e-01, 1.79e-01] [1.52e-01, 7.62e-01, 1.79e-01] [1.90e+00]
1000 [2.71e-03, 1.02e-01, 1.15e-02] [2.18e-03, 1.02e-01, 1.15e-02] [2.80e+00]
2000 [1.33e-03, 3.78e-02, 2.95e-03] [1.12e-03, 3.78e-02, 2.95e-03] [6.16e+00]
3000 [6.58e-04, 2.84e-03, 5.18e-06] [5.12e-04, 2.84e-03, 5.18e-06] [1.02e+01]
4000 [1.45e-04, 1.38e-03, 3.26e-05] [1.10e-04, 1.38e-03, 3.26e-05] [1.07e+01]
Best model at step 4000:
train loss: 1.55e-03
test loss: 1.52e-03
test metric: [1.07e+01]
'train' took 6.077505 s
6 it number
learning rate: 9.8e-03
num_dense_layers: 5
num_dense_nodes: 49
activation: sin
Compiling model...
'compile' took 0.000124 s
Training model...
Step Train loss Test loss Test metric
0 [2.33e-02, 1.04e+00, 4.76e-01] [2.14e-02, 1.04e+00, 4.76e-01] [1.77e+00]
1000 [1.23e-02, 8.18e-02, 3.47e-02] [1.24e-02, 8.18e-02, 3.47e-02] [3.28e+00]
2000 [1.28e-02, 4.04e-02, 1.56e-02] [1.31e-02, 4.04e-02, 1.56e-02] [4.77e+00]
3000 [3.16e-04, 1.88e-02, 1.55e-03] [2.91e-04, 1.88e-02, 1.55e-03] [6.23e+00]
4000 [1.56e-04, 5.74e-03, 3.08e-04] [1.42e-04, 5.74e-03, 3.08e-04] [7.63e+00]
Best model at step 4000:
train loss: 6.21e-03
test loss: 6.19e-03
test metric: [7.63e+00]
'train' took 13.425033 s
7 it number
learning rate: 1.2e-03
num_dense_layers: 3
num_dense_nodes: 29
activation: tanh
Compiling model...
'compile' took 0.000123 s
Training model...
Step Train loss Test loss Test metric
0 [1.13e-02, 6.81e-01, 9.73e-02] [1.10e-02, 6.81e-01, 9.73e-02] [1.56e+00]
1000 [1.40e-03, 2.39e-03, 3.65e-04] [1.50e-03, 2.39e-03, 3.65e-04] [9.56e+00]
2000 [1.30e-04, 9.48e-04, 7.82e-05] [6.60e-03, 9.48e-04, 7.82e-05] [1.03e+01]
3000 [4.26e-04, 6.30e-04, 3.66e-04] [1.09e-02, 6.30e-04, 3.66e-04] [1.05e+01]
4000 [3.74e-03, 1.10e-03, 9.50e-04] [1.17e-02, 1.10e-03, 9.50e-04] [1.06e+01]
Best model at step 2000:
train loss: 1.16e-03
test loss: 7.63e-03
test metric: [1.03e+01]
'train' took 9.438914 s
8 it number
learning rate: 1.8e-03
num_dense_layers: 3
num_dense_nodes: 31
activation: tanh
Compiling model...
'compile' took 0.000108 s
Training model...
Step Train loss Test loss Test metric
0 [1.56e-04, 3.83e-01, 1.47e-02] [1.23e-04, 3.83e-01, 1.47e-02] [1.40e+00]
1000 [3.06e-04, 1.72e-03, 7.34e-05] [1.31e-02, 1.72e-03, 7.34e-05] [1.01e+01]
2000 [1.84e-04, 1.12e-03, 8.03e-05] [9.00e-03, 1.12e-03, 8.03e-05] [1.04e+01]
3000 [4.03e-04, 7.68e-04, 1.86e-04] [9.62e-03, 7.68e-04, 1.86e-04] [1.06e+01]
4000 [8.94e-05, 5.43e-04, 4.65e-05] [1.14e-02, 5.43e-04, 4.65e-05] [1.07e+01]
Best model at step 4000:
train loss: 6.79e-04
test loss: 1.20e-02
test metric: [1.07e+01]
'train' took 9.431659 s
9 it number
learning rate: 1.3e-04
num_dense_layers: 4
num_dense_nodes: 26
activation: sigmoid
Compiling model...
'compile' took 0.000131 s
Training model...
Step Train loss Test loss Test metric
0 [2.27e+00, 6.11e-01, 1.64e-02] [2.27e+00, 6.11e-01, 1.64e-02] [2.52e+00]
1000 [7.87e-07, 2.08e-01, 8.00e-02] [7.44e-07, 2.08e-01, 8.00e-02] [1.45e+00]
2000 [6.23e-07, 2.07e-01, 7.92e-02] [5.78e-07, 2.07e-01, 7.92e-02] [1.45e+00]
3000 [1.36e-04, 2.01e-01, 7.55e-02] [1.27e-04, 2.01e-01, 7.55e-02] [1.43e+00]
4000 [1.91e-03, 1.82e-01, 7.25e-02] [1.56e-03, 1.82e-01, 7.25e-02] [1.40e+00]
Best model at step 4000:
train loss: 2.57e-01
test loss: 2.56e-01
test metric: [1.40e+00]
'train' took 11.026168 s
10 it number
learning rate: 7.7e-04
num_dense_layers: 5
num_dense_nodes: 34
activation: sin
Compiling model...
'compile' took 0.000115 s
Training model...
Step Train loss Test loss Test metric
0 [5.28e-02, 4.02e-01, 7.13e-03] [4.34e-02, 4.02e-01, 7.13e-03] [1.59e+00]
1000 [5.43e-04, 1.70e-03, 6.53e-06] [4.17e-04, 1.70e-03, 6.53e-06] [1.04e+01]
2000 [9.60e-05, 6.60e-04, 2.51e-05] [1.01e-04, 6.60e-04, 2.51e-05] [1.09e+01]
3000 [3.46e-05, 4.73e-04, 2.44e-05] [4.67e-05, 4.73e-04, 2.44e-05] [1.10e+01]
4000 [1.33e-04, 4.08e-04, 2.46e-05] [1.59e-04, 4.08e-04, 2.46e-05] [1.11e+01]
Best model at step 3000:
train loss: 5.32e-04
test loss: 5.44e-04
test metric: [1.10e+01]
'train' took 12.797166 s
11 it number
learning rate: 4.8e-02
num_dense_layers: 1
num_dense_nodes: 5
activation: sin
Compiling model...
'compile' took 0.000124 s
Training model...
Step Train loss Test loss Test metric
0 [1.52e-02, 3.54e-01, 4.04e-02] [1.24e-02, 3.54e-01, 4.04e-02] [1.22e+00]
1000 [5.06e-04, 1.39e-03, 4.19e-05] [4.56e-04, 1.39e-03, 4.19e-05] [1.09e+01]
2000 [2.06e-02, 3.06e-03, 1.99e-03] [2.02e-02, 3.06e-03, 1.99e-03] [1.11e+01]
3000 [2.92e-05, 3.75e-04, 1.28e-05] [2.35e-05, 3.75e-04, 1.28e-05] [1.09e+01]
4000 [7.49e-05, 7.66e-05, 7.82e-05] [7.79e-05, 7.66e-05, 7.82e-05] [1.12e+01]
Best model at step 4000:
train loss: 2.30e-04
test loss: 2.33e-04
test metric: [1.12e+01]
'train' took 6.010061 s
12 it number
learning rate: 2.4e-04
num_dense_layers: 6
num_dense_nodes: 6
activation: sin
Compiling model...
'compile' took 0.000178 s
Training model...
Step Train loss Test loss Test metric
0 [1.10e-01, 2.66e-01, 1.29e-01] [1.06e-01, 2.66e-01, 1.29e-01] [1.22e+00]
1000 [8.82e-03, 1.60e-01, 6.02e-02] [7.93e-03, 1.60e-01, 6.02e-02] [1.61e+00]
2000 [3.15e-03, 8.19e-02, 7.82e-03] [1.93e-03, 8.19e-02, 7.82e-03] [3.78e+00]
3000 [2.52e-03, 2.85e-02, 2.49e-03] [4.10e-03, 2.85e-02, 2.49e-03] [6.72e+00]
4000 [2.67e-03, 1.22e-02, 8.99e-04] [2.73e-03, 1.22e-02, 8.99e-04] [8.28e+00]
Best model at step 4000:
train loss: 1.58e-02
test loss: 1.58e-02
test metric: [8.28e+00]
'train' took 13.344941 s
13 it number
learning rate: 1.2e-04
num_dense_layers: 1
num_dense_nodes: 49
activation: tanh
Compiling model...
'compile' took 0.000135 s
Training model...
Step Train loss Test loss Test metric
0 [1.48e-02, 5.10e-01, 9.98e-03] [1.21e-02, 5.10e-01, 9.98e-03] [1.38e+00]
1000 [4.68e-06, 1.90e-01, 7.80e-02] [4.14e-06, 1.90e-01, 7.80e-02] [1.41e+00]
2000 [2.04e-05, 1.88e-01, 7.72e-02] [1.75e-05, 1.88e-01, 7.72e-02] [1.42e+00]
3000 [3.46e-04, 1.81e-01, 7.36e-02] [2.89e-04, 1.81e-01, 7.36e-02] [1.47e+00]
4000 [7.71e-03, 1.58e-01, 5.82e-02] [6.50e-03, 1.58e-01, 5.82e-02] [1.64e+00]
Best model at step 4000:
train loss: 2.24e-01
test loss: 2.23e-01
test metric: [1.64e+00]
'train' took 6.112236 s
14 it number
learning rate: 5.0e-02
num_dense_layers: 6
num_dense_nodes: 50
activation: sin
Compiling model...
'compile' took 0.000160 s
Training model...
Step Train loss Test loss Test metric
0 [1.19e-02, 4.35e-01, 8.88e-04] [9.82e-03, 4.35e-01, 8.88e-04] [1.47e+00]
1000 [4.16e+08, 1.32e+00, 1.35e-01] [9.42e+18, 1.32e+00, 1.35e-01] [9.99e+00]
2000 [5.94e+10, 1.32e+00, 1.35e-01] [9.42e+18, 1.32e+00, 1.35e-01] [9.99e+00]
3000 [6.80e+10, 1.32e+00, 1.59e-01] [1.12e+19, 1.32e+00, 1.59e-01] [9.97e+00]
4000 [1.42e+11, 1.31e+00, 8.90e-02] [1.40e+19, 1.31e+00, 8.90e-02] [9.65e+00]
Best model at step 0:
train loss: 4.48e-01
test loss: 4.46e-01
test metric: [1.47e+00]
'train' took 15.043787 s
Reentrenamiento con la mejor configuración#
Una vez terminada la búsqueda, se vuelve a crear la PINN usando los hiperparámetros óptimos y se la entrena otra vez. Esto permite disponer del modelo final para hacer predicciones, comparar con la solución exacta y analizar visualmente el resultado.
print("Best params:", best_params)
print("Best score:", best_score)
modelo, data = create_model(best_params)
error, losshistory, train_state = train_model(modelo, best_params, iterations=10000)
Best params: [0.047782941634808326, 1, 5, 'sin']
Best score: 0.00023269429
Compiling model...
'compile' took 0.000121 s
Training model...
Step Train loss Test loss Test metric
0 [6.09e-03, 6.21e-01, 6.52e-02] [5.45e-03, 6.21e-01, 6.52e-02] [1.47e+00]
1000 [5.45e-08, 2.63e-04, 1.25e-05] [3.75e-08, 2.63e-04, 1.25e-05] [1.08e+01]
2000 [3.56e-08, 3.29e-05, 1.63e-06] [3.47e-08, 3.29e-05, 1.63e-06] [1.08e+01]
3000 [3.22e-08, 4.00e-06, 2.09e-07] [3.64e-08, 4.00e-06, 2.09e-07] [1.08e+01]
4000 [2.58e-08, 7.92e-07, 3.25e-08] [2.91e-08, 7.92e-07, 3.25e-08] [1.09e+01]
5000 [7.18e-07, 2.53e-06, 4.14e-06] [8.09e-07, 2.53e-06, 4.14e-06] [1.09e+01]
6000 [1.21e-03, 3.90e-05, 6.87e-05] [1.36e-03, 3.90e-05, 6.87e-05] [1.09e+01]
7000 [2.79e-08, 3.59e-07, 6.39e-09] [3.14e-08, 3.59e-07, 6.39e-09] [1.09e+01]
8000 [1.70e-07, 2.75e-07, 5.85e-09] [1.91e-07, 2.75e-07, 5.85e-09] [1.09e+01]
9000 [6.56e-05, 1.25e-06, 4.03e-06] [7.39e-05, 1.25e-06, 4.03e-06] [1.09e+01]
10000 [8.43e-10, 1.56e-08, 5.53e-10] [8.90e-10, 1.56e-08, 5.53e-10] [1.09e+01]
Best model at step 10000:
train loss: 1.70e-08
test loss: 1.71e-08
test metric: [1.09e+01]
'train' took 14.839695 s
Evaluación sobre puntos del dominio#
Acá se generan puntos del intervalo para inspeccionar la solución aprendida. Se calculan:
las predicciones en los puntos observados,
las predicciones en una malla más densa del dominio,
los valores exactos para comparar.
Con esto se puede verificar si la red reproduce correctamente la forma esperada de la solución de la PDE.
x = geom.uniform_points(50, True)
X_domain = data.train_x[:data.num_domain]
y_pred_train = modelo.predict(x_train)
y_exact_train = sol_exacta(x_train)
y_pred = modelo.predict(x)
y_exact = sol_exacta(x)
Visualización de la solución y del proceso de optimización#
Las últimas celdas muestran dos tipos de resultados:
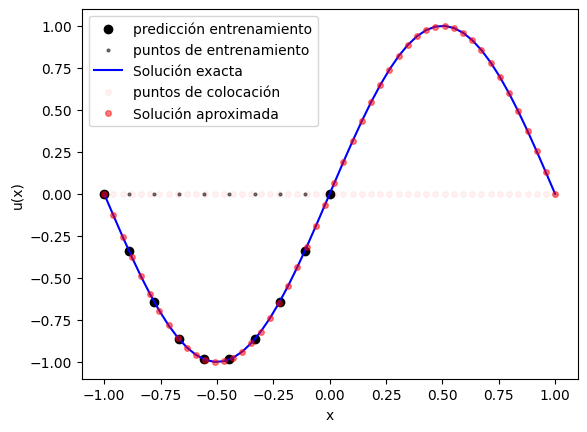
la comparación entre solución exacta, predicción de la PINN y puntos usados en el entrenamiento;
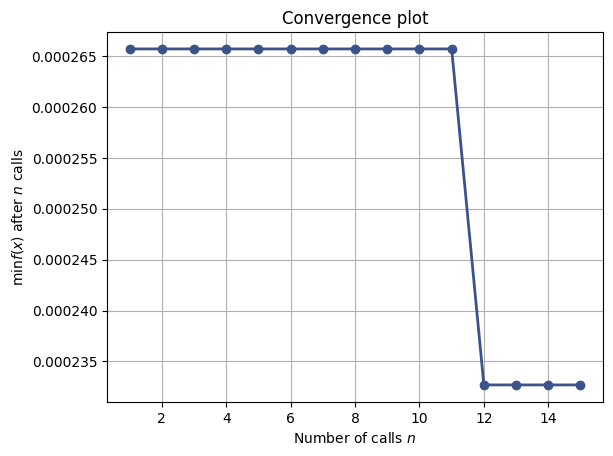
la evolución de la optimización bayesiana mediante
plot_convergenceyplot_objective.
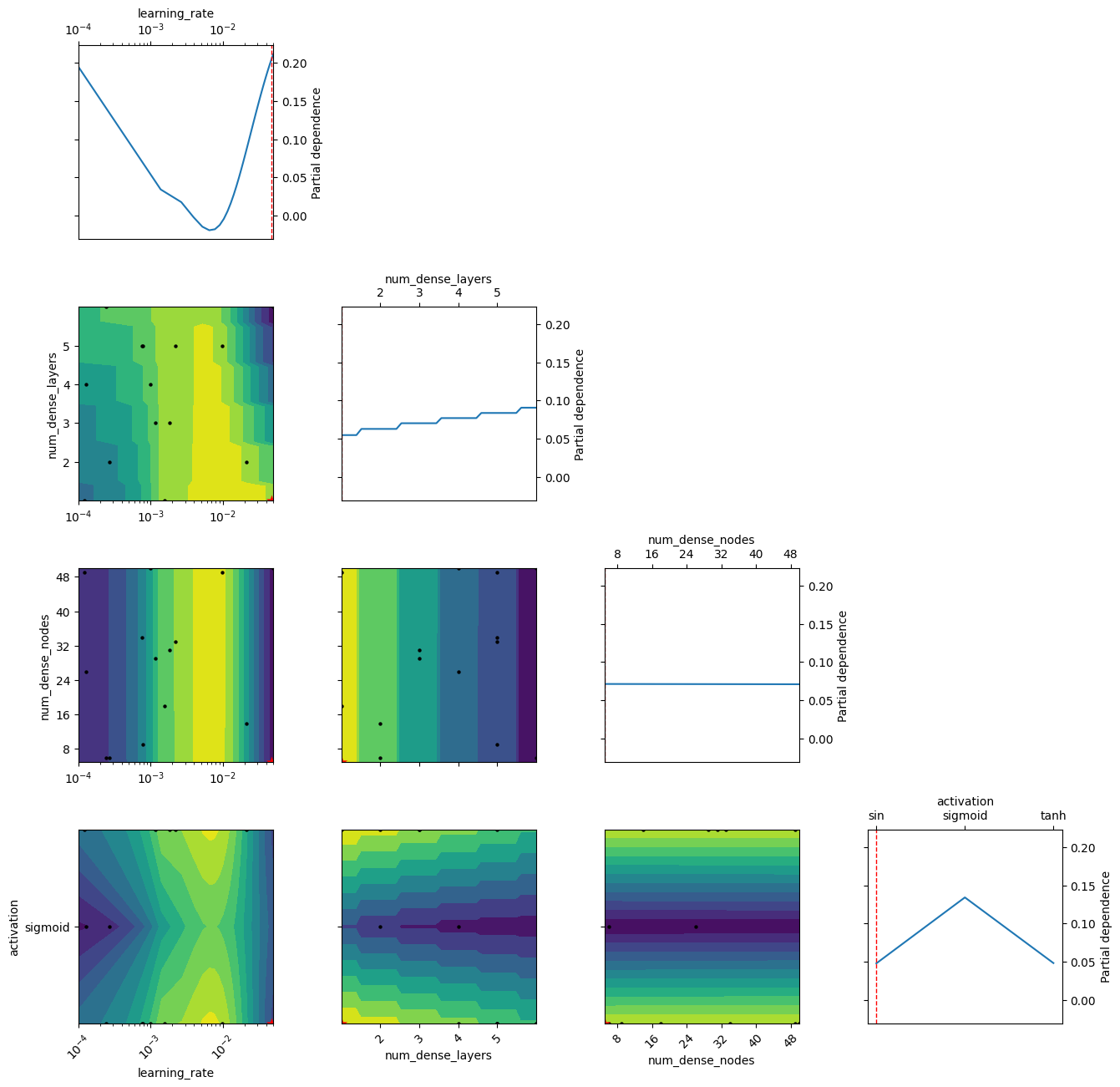
La primera figura sirve para validar la aproximación física. Las otras ayudan a entender si la búsqueda de hiperparámetros convergió y cómo influyó cada variable sobre el desempeño del modelo.
plt.figure()
plt.scatter(x_train, y_pred_train[:,0], color='black', label="predicción entrenamiento")
plt.scatter(x_train, np.zeros_like(x_train), color='black', label="puntos de entrenamiento", alpha=0.5, s=4)
plt.plot(x, y_exact, 'b', label="Solución exacta")
plt.plot(x, np.zeros_like(x), 'or', label="puntos de colocación", alpha=0.05, ms=4)
plt.plot(x, y_pred[:,0], 'or', label="Solución aproximada", alpha=0.5, ms=4)
plt.xlabel("x")
plt.ylabel("u(x)")
plt.legend()
plt.show()
plot_convergence(search_result)
plt.show()
plot_objective(search_result, show_points=True, size=3.8)
plt.show()
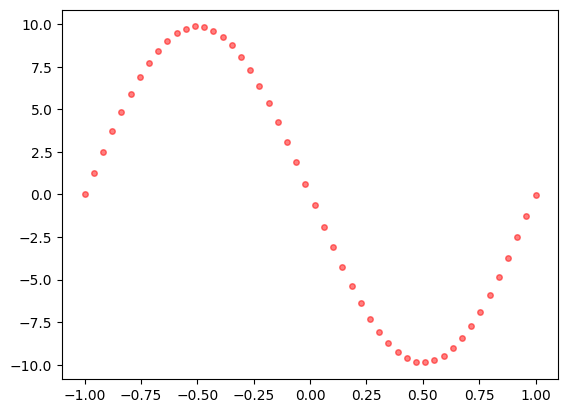
Vemos la estimación del campo de fuerza
plt.plot(x, y_pred[:,1], 'or', label="Solución aproximada", alpha=0.5, ms=4)
plt.show()